By Mathieu Boudreau
The August 2020 MRM Highlights Reproducible Research Insights interview is with Steven Whitaker, Jon-Fredrik Nielsen and Jeff Fessler, researchers at the University of Michigan, and authors of a paper entitled “Myelin water fraction estimation using small-tip fast recovery MRI”. This paper was chosen as the MRM Highlights pick of the month because it reports good reproducible research practices. In particular, in addition to sharing code used in their work, the authors created and shared scripts that reproduce every figure from their paper. To learn more about Steven, Jon-Fredrik, Jeff and their research, check out our recent interview with them.
General questions
1. Why did you choose to share your code/data?
We have benefitted from open source software in the past, so sharing code is one way of saying thanks. Additionally, it is impossible, in a paper, to completely describe the exact methods used; there’s just not enough space, and even if there was, the code itself often describes the methods in a more concise (and less ambiguous) manner. In other words, the Methods section of a paper essentially provides a summary or highlights of the methods, but if you want to know exactly how the data were processed then you need to look at the code. Finally, sharing code allows others to more easily learn about and build on the research we have done.
I (Steven) am a little over halfway through my PhD program, and so I look at this aspect through the lens of a new researcher, just getting started on their first project. When I was first learning about small-tip fast recovery (STFR, the type of scan used in our paper), I wrote my own code in order to reproduce the figures from STFR papers. It served as a learning exercise. But there were one or two figures that I couldn’t reproduce on my own, so it was helpful to have access to the authors’ code to help fill in the gaps in my understanding. (Incidentally, I found a typo in one of the papers, so their code was correct, but the Methods section was not!)
We decided to share the raw data as well, in order to facilitate future research where researchers might need access to more data than they would be able to obtain on their own.
2. Do you think you’ll share code/data again in future publications?
Definitely!
3. At what stage did you decide to share your code/data? Is there anything you wish you had known beforehand, or done sooner?
From the very outset, it was always our intention to make our code publicly available once our paper was accepted for publication. I don’t remember exactly when we decided to make the data available, it just seemed like a good idea so we went with it.
4. How do you think we might encourage researchers in the MRI community to contribute more open source code along with their research papers?
It could be an idea to include a reminder about source code and data in the submission system, as a means of highlighting this aspect when people first submit their papers to MRM. Not as a requirement, but as a way of sending out a strong message that this is encouraged. Perhaps the system could even ask “why not?” if authors indicate that they don’t plan to share code/data. (There can, of course, be legitimate reasons why not – IRB issues with human data for example.)
Questions about the specific reproducible research habit
1. What practical advice do you have for people who would like to write code that creates reproducible figures to be shared along with their paper?
First, start the research with a reproducible mindset, particularly when writing the first line of code. If the code is written clearly in the first place, this will probably help the research to progress better and it makes it far easier to share the finished code at the end. In contrast, taking hastily thrown together code and trying to turn it into code suitable for reproducible research later on will undoubtedly prove laborious, and might not even allow proper reproduction of the results.
Second, separate number-crunching from plotting. It can take a while to get the plot to look just right, and you don’t want to have to keep running your code just to tweak plotting parameters, as this can take minutes or even hours. Run the code once, save the results, and have the plotting code load the results from disk.
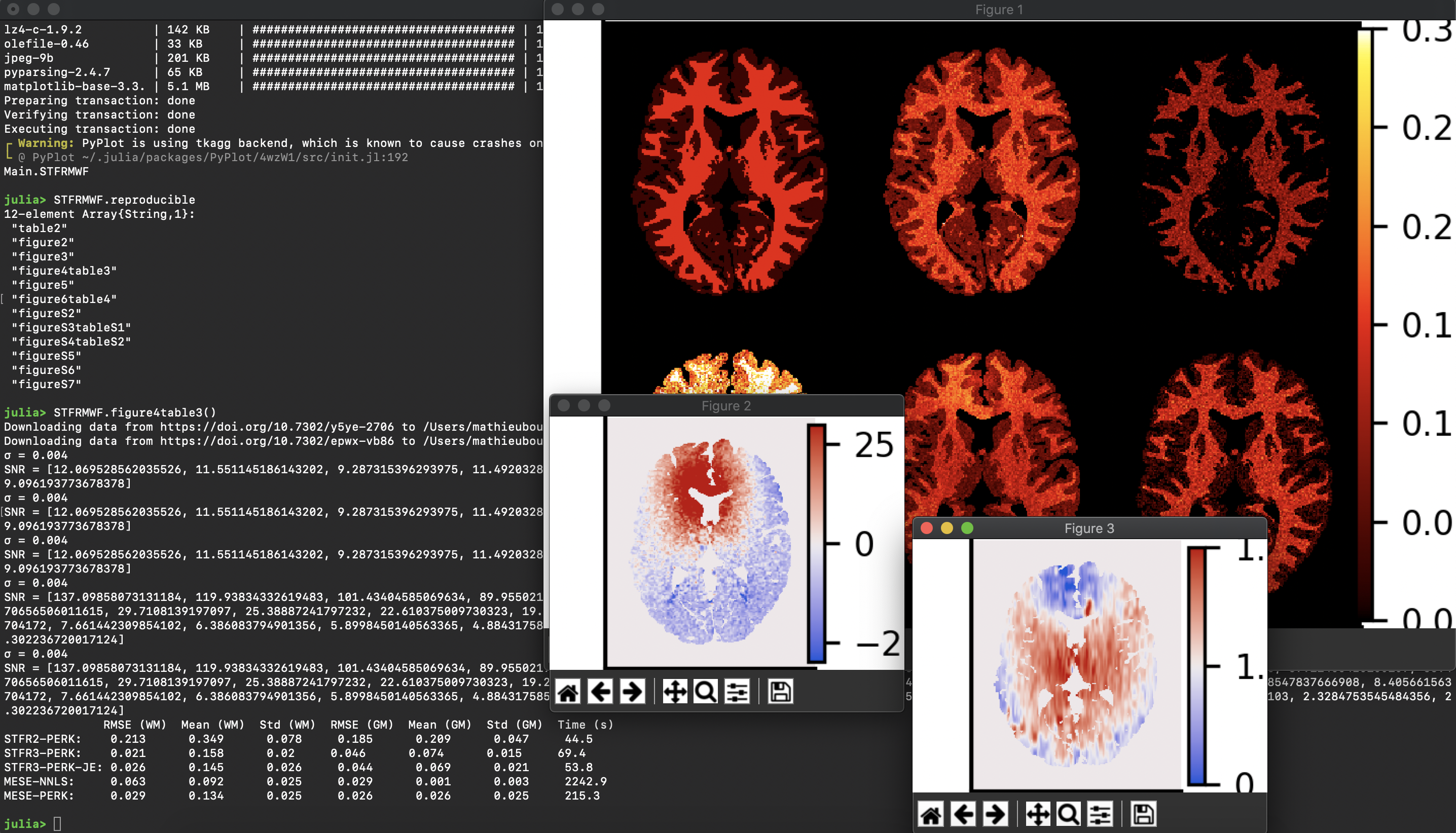
2. Did you encounter any challenges or hurdles while developing the scripts/code that reproduce your figures?
Initially the data repository at the University of Michigan, which is called “DeepBlue”, had a digital object identifier (DOI) for the top-level repository, but we wanted an individual DOI for each data file so as to be able to provide the code in a very clean way that links it directly to the exact files in the repository. Fortunately, the DeepBlue staff were happy to work with us to provide a DOI for each data file. As a result, the final version is likely the most reproducible work ever to come out of the Fessler group at Michigan.
3. Why did you choose Julia as a programming language for this project?

There are many benefits to be had from using Julia. It is an interactive language (dynamically typed), like MATLAB and Python, and this makes it convenient for code development. But it is built on a compiler “under the hood”, so you get execution speeds like you do with compiled code. Julia has very advanced package management features that are closely integrated with git so that when we release our code, it is associated with specific versions of all of the libraries that we used; in this way, later on, when someone uses our code to reproduce the results, it should use exactly the same library versions that we used for the paper even though those libraries will have evolved in the meantime. Julia also supports distributed computing, GPU acceleration and deep learning libraries, though we did not use those capabilities in this work. There is now a Julia version of the Michigan Image Reconstruction Toolbox (MIRT) that provides tools for MRI research, like NUFFT operations and regularizers. The way Julia is integrated with GitHub makes it convenient to write test routines for Julia code, and MIRT has nearly 100% code coverage for its tests. The “parameter estimation via regression with kernels” (PERK) library developed in Julia for this work has 100% code coverage, which can help give potential users confidence in the code.
Julia is also free and open source, while MATLAB is not. This is an especially important consideration if one uses MATLAB toolboxes beyond a default installation of MATLAB, because even if someone else has access to MATLAB through their institution, they might not have access to the specific toolboxes you used, which limits the reach of your open source code.
4. Are there any other reproducible research habits that you haven’t used for this paper that you might be interested in trying in the future?
Both MIRT.jl and PERK.jl have “docstrings” that internally document each function, but neither has a separate “user manual”, so there is always room for improvement in documentation.
Jupyter Notebooks are a nice way to interleave code and results with an explanation of what is going on. They can be useful for demonstrating how code is intended to be used.




