By Mathieu Boudreau
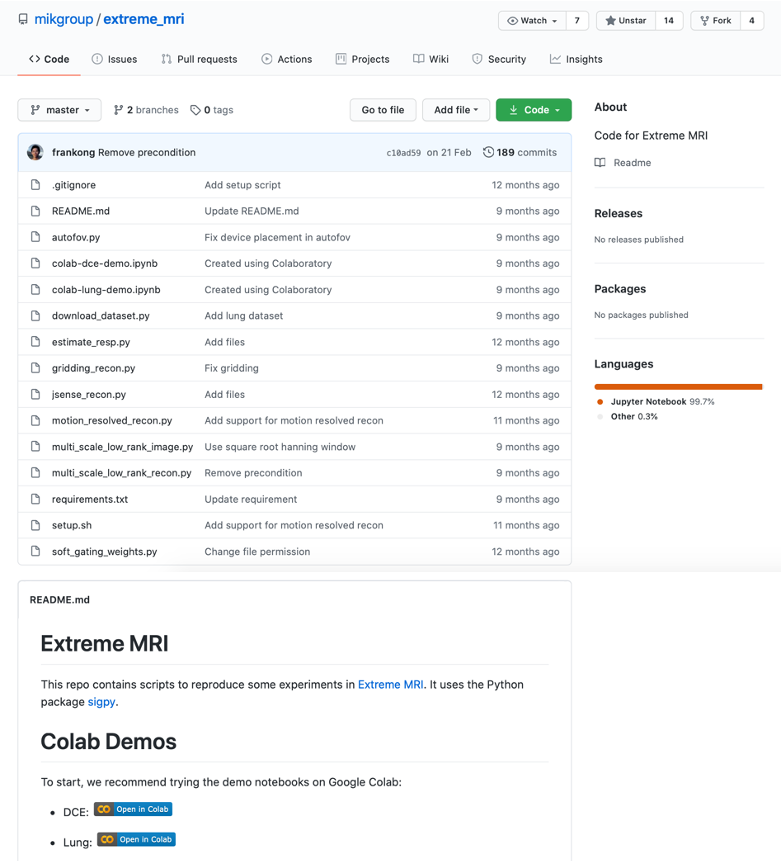
The October 2020 MRM Highlights Reproducible Research Insights interview is with Frank Ong and Miki Lustig, researchers at Stanford and Berkeley respectively, and authors of a paper entitled “Extreme MRI: Large‐scale volumetric dynamic imaging from continuous non‐gated acquisitions”. This paper was chosen as the MRM Highlights pick of the month because it reports good reproducible research practices. In particular, in addition to sharing code used in their work, the authors also shared a demo of their work in a Google Colab notebook. To learn more about Frank and Miki and their research, check out our recent interview with them.
General questions
1. Why did you choose to share your code/data?
[Miki] As a graduate student, I was working with Profs. John Pauly and David Donoho. John Pauly was one of the first researchers in MRI to share their code freely, implementing a suite of tools for RF pulse design (RF Tools). David Donoho was already releasing code for every single one of his papers, so the culture and environment promoted sharing. In particular I was inspired by the paper “WaveLab and Reproducible Research” by Buckheit and Donoho, in which they quote Prof. Claerbout – a pioneer in reproducible research: “An article about computational science in a scientific publication is not the scholarship itself, it is merely advertising of the scholarship. The actual scholarship is the complete software development environment and the complete set of instructions which generated the figures”. I followed the paths of my mentors and passed this mindset on to my mentees. Frank has definitely taken this to another level, with the release of a SigPy — a complete signal processing and optimization package. Over the years, we have found that papers that share code and data end up being much more influential than those that don’t. I recall the fierce resistance to code sharing at the first Sedona workshop in 2007. The winds have definitely changed, and you felt that at the 2020 Sedona meeting. This trend will likely continue and hopefully become a requirement for publication.
2. What is your lab or institutional policy on sharing code/data?
The sharing culture is embedded within UC Berkeley. In fact, the Berkeley Software Distribution (BSD) license is an extremely permissive free software license with minimal restrictions on use and distribution. This culture is also embedded in the lab.
When lab members submit a paper, they try to make it as easy as possible to reproduce. That means releasing code and data, and also creating demos and documentations. Miki’s compressed sensing and SPIRiT toolboxes set a great example for the lab.
There’s also a culture to create general software libraries with permissive licenses, like BSD when possible, so that other groups can leverage these sources to develop their own projects. Martin Uecker, who is an alumnus of the group, and now faculty at the University Medical Center Göttingen, pioneered this with BART, which is now used in multiple research groups and institutions.
3. At what stage did you decide to share your code/data for this project? If it was late in the project, is there anything you wish you had known or done sooner? If it was early on, what did you do in preparation to do so?
For this project, we always intended that the code and data would be shared. So, to keep track of what was being done, we followed common software development practices for version controlling, documentations and unit tests. These practices proved particularly helpful for this project because of the sheer complexity of the overall pipeline. In the process of doing so, we also created a package called SigPy, which provides a general framework in Python for iterative reconstruction using multi CPU and GPUs. This allowed us to separate code that can be leveraged for other projects and code that is specific to this project.
4. How do you think we might encourage researchers in the MRI community to contribute more open-source code along with their research papers?
Showcasing works with open-source code is a great way to encourage researchers. In this sense, MRM Highlights has really done a great job with these interviews. The hands-on software sessions in ISMRM have also attracted a lot of attention in recent years. We think doing more of these will further encourage researchers to contribute open-source code.
Questions about specific reproducible research habits

1. What advice do you have for people who would like to use Jupyter Notebooks and Google Colab?
Jupyter notebooks are great for demos and interactive data processing. Here are some tricks we find useful:
- %matplotlib notebook in the beginning of a notebook enables interactive matplotlib plotting.
- %timeit and %%timeit can time a function in line and cell mode respectively.
- tqdm is a great package that allows you to create progress bars with custom information.
- ! allows you to run shell commands.
Google Colab is an amazing resource for creating demos that use GPUs. It requires little setup on the user side and allows anyone to access powerful GPUs for free. But Google Colab has a slightly different interface to Jupyter Notebook. So, if you don’t need GPUs for your demo, we recommend using MyBinder instead.
2. Can you share some resources to help readers get started with these tools?
There are now a lot of notebooks on MRI data processing available online. A great way to get started is to go through them. Here are some examples:
SigPy reconstruction and pulse design tutorials using Jupyter Notebook:
- https://github.com/mikgroup/sigpy-mri-tutorial
- https://github.com/jonbmartin/open-source-pulse-design
BART tutorial using Jupyter notebooks:
Dictionary learning tutorial using Juypter notebooks:
Deep learning tutorial using Jupyter and Colab notebooks:
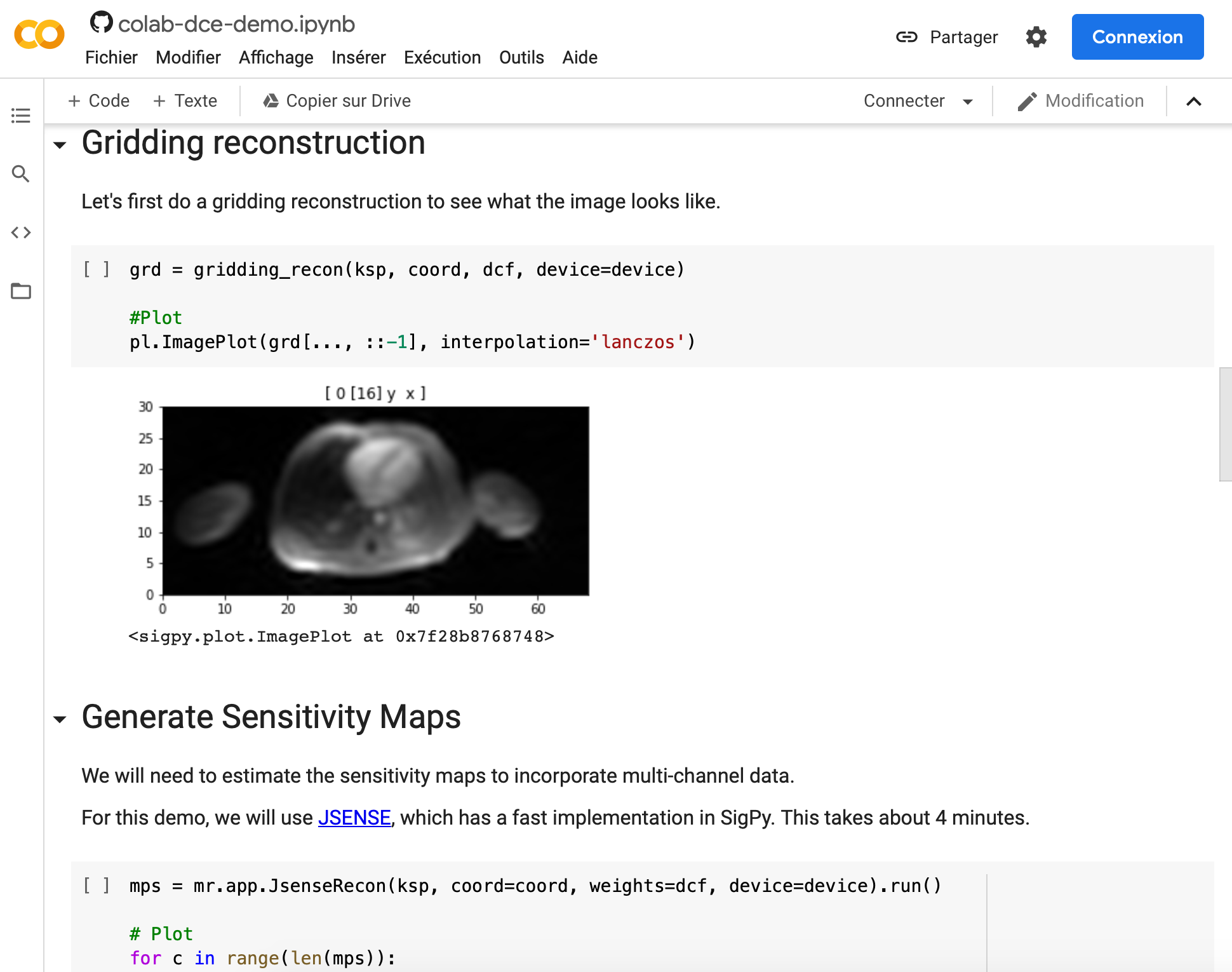
3. Did you encounter any challenges or hurdles during the process of developing open-source tools and demos requiring GPU computing?
In the past few years, many tools have been developed that make it really easy to develop open-source packages and demos requiring GPU computing. Installing GPU drivers and packages used to be difficult. A lot of tools needed to be created from scratch. Now, there are a lot of open-source GPU packages available, such as CuPy (a GPU version of NumPy) and of course Tensorflow and PyTorch for deep neural networks. They provide an easy way to leverage GPU for numerical computation and they are all open source. And Google Colab makes it really easy to create demos that use GPUs because there’s no installation required on the user side.
4. Are there any other reproducible research habits that you haven’t used for this paper but might be interested in trying in the future?
We’d like to explore Zenodo more for data hosting. The ability to cite individual datasets and keep track of them is really powerful. And Zenodo has expanded its data limits over the years. We’ve uploaded several datasets for the demo of this project. But moving forward, it might be interesting to use Zenodo during method development, too.




