By Mathieu Boudreau
The June 2021 MRM Highlights Reproducible Research Insights interview is with Ziwei Zhao and Yongwan Lim (co-first authors) and Krishna S. Nayak, researchers at the University of Southern California in Los Angeles, California. Their paper is entitled “Improved 3D real-time MRI of speech production”. It was chosen not only because the authors share code and data with their paper, but also because their code repository is well documented and they used the BART toolbox in their implementations. To learn more about this team and their research, check out our recent interview with them.
To discuss this Q&A, please visit our Discourse forum.
General questions
1. Why did you choose to share your code/data?
So many reasons! First, we want the work to be accessible and have the maximum possible impact. We believe that sharing the software implementation and sample data are complementary to the manuscript itself, and this practice increases the likelihood that the techniques will be utilized by readers in their own work. Second, we want all readers to understand the guts of the technique, and this is best done by presenting material in multiple ways. For most readers, practicing with provided code and data can expedite their understanding of the work. Third, we wanted to “pay it forward”. We benefited from the developers and maintainers of BART and would therefore like to continue their tradition. We provided a complete example that uses BART for the constrained reconstruction of dynamic 3D datasets. Finally, we are all individually supportive of the growing culture of reproducible research within our ISMRM community and are happy to be a part of it. We hope that readers take advantage of our shared code and data and have fun with them.
2. What is your lab or institutional policy on code/data sharing?
We have a lab policy to share our project-related code on private GitHub repositories that are accessible to all current lab members. This includes work in the development stage a.k.a. “sandbox”. This habit has allowed us to understand each other’s work better and to identify issues and bugs, to improve our coding and documentation style, and thus to make it easier to reuse code written by others. Neither our lab nor our institution has a formal policy on external code/data sharing. However, all of our lab members are enthusiastic about this practice.
3. At what stage did you decide to share your code/data? Is there anything you wish you had known or done sooner?
Our code and data were shared within our lab, under privacy protection, from the onset of this project. We decided to publish our code and data while we were preparing the manuscript, in other words at a stage when most of the work had been done. It would be great to have shared a comprehensive reconstruction package that caters not only for 3D but also for 2D speech protocols. We may revisit this aspect when time permits.
4. How do you think we might encourage researchers in the MRI community to contribute more open-source code along with their research papers?

Giving such papers priority in the selection process for honors, such as MRM Highlights, is a great way to encourage the practice. Other than that, we just have to spread the word about how this increases impact. We have all seen great examples of published open-source code and datasets, having an incredible impact on the pace of research in our ISMRM community, and making our research more inclusive. For example, the impact of the BART reconstruction toolbox has been immense, and has broken down barriers for beginners. The reproducible research sessions at recent years’ ISMRM meetings appear to be attracting a growing audience. Their hands-on demos and open discussions are very helpful, and are increasing awareness of open-source research.
Questions about the specific reproducible research habit
1. What advice do you have for people who would like to create well-documented code repositories?
One general piece of advice would be to identify and closely examine examples of well-documented and well-written code repositories. Then apply, in your own code repositories, any best practices and style that you admired in the examples. This has worked within our laboratory, and it is how we have improved our coding skills. We summarize some specific tips below.
Preparation
- It’s a great idea to ask yourself what information you want to convey to the readers. This should be closely related to the main points of the paper. For example, improvements of the results or accelerated computing times should be reflected in the implementation. It is also helpful to include the scripts that generate critical figures from the paper.
- A good README file will help readers understand the content (functions, scripts, datasets) and will elaborate on the work.
Coding habits
- Comment your code liberally. For example, state the purpose of each section; state the relevant formula if useful, comment on the important unit, etc.
- Follow simple, self-explanatory, and consistent naming conventions for all variables, functions, and so on. Cultivate your coding habits in such a way as to align with established programming style.
Feedback
- Ask labmates to review your work and give constructive suggestions.
- Solicit a test audience, ideally ones who are not familiar with your work, in order to run beta tests, and ask them for feedback on readability and ease-of-use.
2. What questions did you ask yourselves while you were documenting your code?
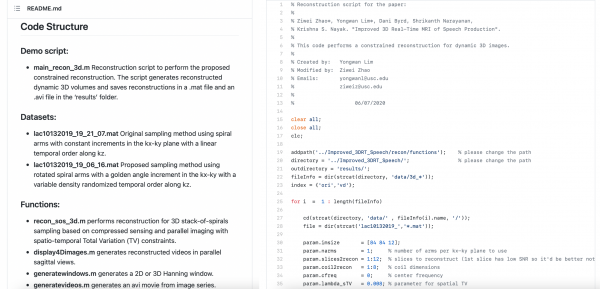
We asked ourselves how we could most efficiently make the code available for usage within our lab and for publishing. The scripts were widely used each time we acquired new data, and were shared with several lab members. To keep everything clean and easy for users to understand and execute, we deliver the limited important parameters in the main function and keep the details of the reconstruction process fixed in separate functions. In addition, we asked ourselves whether the repository would be easy to learn and understand, even for people not familiar with the work, if published.
3. Why did you choose to use an external MRI tool (BART toolbox) instead of developing your implementations yourselves? Can you share some resources to help our readers get started with the BART toolbox?
Yongwan: It was natural for me to implement the code of this work using the BART toolbox. I have used the BART toolbox since 2015. Around that time, our lab was about to start a huge data collection and processing project that would take the next couple of years, and we needed a more efficient and scalable implementation for the reconstruction pipeline. Since then, our lab has used BART in several projects. I also develop my own implementations every now and then, but it was natural for us to use the toolbox for this work. To beginners using the BART toolbox, I would say subscribe to the mailing list. People discuss issues and workarounds with other users and developers in the forum. I have benefitted a lot from it.
Ziwei: I think BART can generalize the reconstruction process. Our lab has tried to develop the in-house implementation as well. However, implementations can differ between different people, and it is hard to keep them consistent across different developers. Besides, the BART toolbox is a popular reconstruction package in the MRI community. It provides most of the essential reconstruction functions. It also provides a simpler way to edit and tune the parameters. For beginners about to use BART, I recommend going through the BART webpage. There, they can find more examples using BART. The installation details can be found here.
4. Are there any other reproducible research habits that you didn’t use for this paper but might be interested in trying in the future?
Quite a lot. We realized that we could improve the code using shell scripts with BART. This will be beneficial for executing the code on the server without relying on MATLAB. Beyond this, there are benefits to be had from using notebooks like Julia, Jupiter etc. to document the code, and provide a nice UI interface. One great example is the repository demo in this year’s ISMRM educational talk ‘Reconstruction of Non-Cartesian Data’. Readers can directly see the impact of parameter tuning by interactively sliding the bars. It would be fun to use similar formats in our future repositories.




