By Mathieu Boudreau
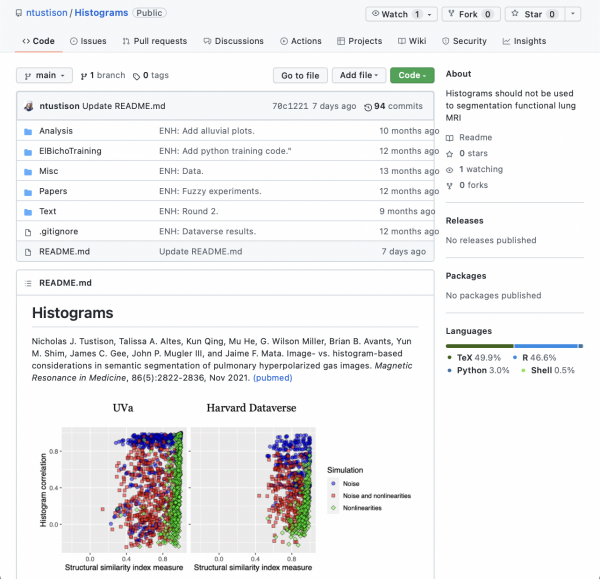
This MRM Highlights Reproducible Research Insights interview is with Nick Tustison, a researcher at the University of Virginia. The paper is entitled “Image- versus histogram-based considerations in semantic segmentation of pulmonary hyperpolarized gas images”, and it was recently discussed by Nick Tustison and Jaime Mata in a Q&A feature. This work was chosen because, in it, the authors demonstrated exemplary reproducible research practices; in particular, they shared the scripts and data required to reproduce every figure published in the paper. To learn more about the work done by Nick Tustison and Jaime Mata, check out our recent interview with them.
To discuss this blog post, please visit our Discourse forum.
General questions
1. Why did you choose to share your code/data?
I did a postdoctoral stint at the University of Pennsylvania, in the PICSL, with Jim Gee who is a longtime supporter of open-source software, having been one of the original developers of the Insight Toolkit (ITK). While at PICSL, I contributed a number of classes to the ITK library while also starting development of the open-source Advanced Normalization Tools (ANTs) software with fellow PICSL alumnus (and co-author) Brian Avants. Following an open-source model for writing code has significantly enhanced the quality and impact of my work, and increased my collaborative opportunities. Sharing the code and data for this particular work is a simple extension of this long history of open-source development. My slightly more cynical attitude towards *not* sharing code, data, etc., is well summarized by a quote Brian once brought to my attention from David Donoho (professor of statistics at Stanford University), giving credit to advice received early in his career from a colleague, “an article about computational results is advertising, not scholarship. The actual scholarship is the full software environment, code and data that produced the results.”
2. What is your lab or institutional policy on code/data sharing?
Being a “lab” of N=1 has its advantages. In my case, I can easily follow my own pro-code/data sharing policies. 🙂 Fortunately, I work with great collaborators who understand the importance and benefits of open-source software. With respect to neuroimaging research at UVA, my principal collaborator, James Stone (Vice Chair of Clinical Research for the Radiology & Medical Imaging Department), has been an important advocate of our open-source developmental work, and this has facilitated institutional support.
3. At what stage did you decide to share your code/data? Is there anything you wish you had known or done sooner?
To help with organization and promote collaboration, I started writing manuscripts and grants a few years ago using a dedicated GitHub repository (one per document). Although I utilize collaborative platforms such as Google Docs and Overleaf, they are somewhat limited in terms of more fully supporting computational reproducibility. I prefer having the tabulated data, statistical analysis scripts, and plotting scripts stored with the text, and this is easily done within a GitHub repository. Simultaneously, it helps you write the document as well as remember little details that are otherwise often forgotten (e.g., tweaks made to generate a particular plot). I have not fully migrated to the strategy of creating a single Rmarkdown file to generate the finished document, for example, but I am typically not far off that approach.
4. How do you think we might encourage researchers in the MRI community to contribute more open-source code along with their research papers?
Personally, I don’t do much open-source proselytizing outside my relatively small sphere of influence. I think certain funding and publishing requirements have helped in a top-down way, but, more importantly, I believe I see evidence of increased grassroots buy-in from the general research community. For example, the growth of pre-publication repositories, like arxiv.org and biorxiv.org, and the growing tendency to explicitly mention source code availability in published materials are, to me, indications that the efforts of early open-source pioneers are beginning to produce the desired results. However, it is also important to note that open source is only one component of the much larger goal of computational reproducibility.
Questions about the specific reproducible research habit
1. What advice do you have for people who would like to contribute to large or established software projects, like the Advanced Normalization Tools (ANTs)?

I would recommend initially exercising the functionality of those software projects which have relevance to one’s personal research interests. That is probably the most direct way of identifying potential contributions and motivating oneself to actually contribute. It should be noted that contributions are not limited to new functionality. For example, Hans Johnson (University of Iowa) is constantly updating ANTs code with ITK developments. Others, like Phil Cook and Jeff Duda (University of Pennsylvania), as well as Gabriel Devenyi (McGill University), contribute by updating code and installation scripts, answering user questions on the forum, and adding new functionality (e.g., Docker containers). We also have R and Python ANTs interfaces with each benefitting from domain expertise for code development and maintenance, specifically the expertise of John Muschelli (Johns Hopkins University) and Nick Cullen (Lund University), respectively. So, although Brian and I have been developing ANTs from the beginning, the entire ANTsX software ecosystem is very much a community effort.
2. What considerations went into ensuring that this software can be used, maintained and/or improved in the long term (on the user or the developer side)?
For this particular work, I wrote quite a bit of code that had more general application potential. So, instead of simply limiting the implementations to the GitHub repository for this paper, I actually ported the more general algorithmic implementations (e.g., fuzzy c-means, histogram warping) to the different libraries within the ANTsX ecosystem, i.e., ANTsR/ANTsPy for image processing and ANTsRNet/ANTsPyNet for deep learning. This has potential benefits not only for my future work, but also for other ANTs users.
3. What motivated your decision to share manuscript documents (drafts, response to reviewers, etc.) openly in your GitHub repository?
As I mentioned previously, I have been using GitHub as a collaborative resource for writing manuscripts for some time. Writing a publication, at least for me, is an important academic exercise and I learn much from the process. In addition to the minutiae that I discover for analyzing and plotting data with particular packages, I benefit significantly from the iterative reviewer interaction. In my opinion, these GitHub repositories become a digital lab book of sorts to which I can return to refresh memories of things I had learned previously. This information could potentially help others, which is why I make the repositories public, although, admittedly, I should probably try to improve the overall organization.
As an illustration of manuscript writing as an important academic experience, during one of our joint projects a few years ago, Brian and I stumbled across an interesting phenomenon which eventually became the topic of a publication in Human Brain Mapping. One of the reviewers, who is very well known and the author of the first paper I ever read as a grad student, affixed his name to the top of his review (which appeared to have been written on an actual typewriter). Although the reviewer wrote approvingly of our work, there was much discussion that was beyond my knowledge at the time (and probably still is). Given the identity of the reviewer, the unusual way in which the review was presented, and the topics discussed, I kept it for sentimental reasons. Unfortunately, however, I didn’t keep the entire record of the interaction, which I now regret.
4. Are there any other reproducible research habits that you didn’t use for this paper but might be interested in trying in the future?
As mentioned previously, I fall short of using Rmarkdown documents as self-contained sources for a given publication. In this sense, the use of Jupyter notebooks is something with which I’d like to become more proficient. Since the ANTsX ecosystem comprises both R and Python libraries, it would really help, in terms of reproducibility, to leverage the tools that have been created for these platforms.




