By Mathieu Boudreau
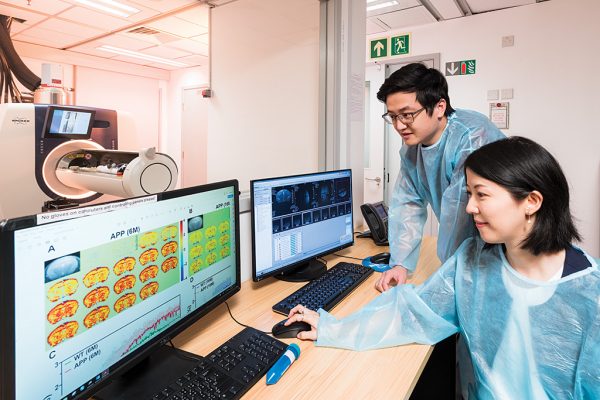
This MRM Highlights Reproducible Research Insights interview is with Jianpan Huang and Kannie WY Chan, researchers at City University of Hong Kong. Their paper is entitled “Deep neural network based CEST and AREX processing: application in imaging a model of Alzheimer’s disease at 3 T”, and we chose to highlight it because the authors demonstrated exemplary reproducible research practices; in particular, they shared code and sample data in addition to a demo and scripts that reproduce some of their figures. To learn more about Jianpan and Kannie and their paper, check out our recent interview with them.
To discuss this blog post, please visit our Discourse forum.
General questions
1. Why did you choose to share your code/data?
There are two main reasons. First of all, we chose to do it for reproducibility purposes. CEST analysis using deep learning is an emerging CEST postprocessing method that is not yet widely used in this setting. Our hope, in sharing our code/data, is that researchers who are working in the CEST field and/or are interested in deep learning-based CEST analysis might be able to use our code directly for processing their own CEST data. The networks in this study were trained on CEST data acquired from mice. Although the trained networks cannot be used directly to perform prediction tasks in other CEST studies, the shared code could easily be applied for training different CEST datasets, such as human CEST data or CEST data acquired using other CEST parameters. This could make it possible to generate the specific networks for prediction. The second reason is to promote the deep learning based CEST/AREX postprocessing method. We believe that this method would have more impact if it were easier for researchers to use the shared code and apply it to their studies. This could provide an avenue for researchers to share their ideas and thus help to improve this approach.
2. What is your lab or institutional policy on code/data sharing?
Currently, our lab does not have a formal policy on code sharing. We encourage members to share code that has been developed in our lab, especially when the related work has reached the publication stage. This is good practice for making code open and transparent, thereby allowing internal or even external researchers to validate the findings and use the methods. As regards data sharing, we share the preclinical animal data among our members for research purposes. Decisions on whether data can be shared to external researchers/repositories are guided by our institutional data policy.
3. At what stage did you decide to share your code/data? Is there anything you wish you had known or done sooner?
We made this decision at the time of submitting the revised manuscript, which is the appropriate stage at which to decide on code/data sharing. With hindsight, it would have been useful to consider the possible issues with code sharing while actually developing the code, so as to reduce the time spent checking and revising it before uploading.
4. How do you think we might encourage researchers in the MRI community to contribute more open-source code along with their research papers?
The MRM Highlights initiative is one way of encouraging this. In addition, I think the journal could perhaps consider flagging published papers that provide ‘open-source code’, also by way of an acknowledgement. Also, if “open-source code” were among the filters that readers can apply when using the Search function, it might be easier to find studies offering this.

Questions about the specific reproducible research habit
1. What advice do you have for people who would like to share their deep learning code/data along with their paper?
It is helpful to share the code used for generating the main figures in the paper, so that readers can easily follow the paper and get started with the code. Furthermore, since it is practically impossible to train two identical networks, even if the training dataset and hyper parameters are the same, it is always advisable to save the networks, especially those used to generate the results presented in papers. When sharing deep learning code/data, it is also useful to add descriptions about how to implement the code (prerequisites and procedures) and, for the benefit of users, to suggest possible errors/bugs.
2. What questions did you ask yourselves while you were developing the code that would eventually be shared?
One aspect considered was how to make the code easy to use, which meant asking ourselves a series of questions. What kind of data format is more convenient if readers want to try this method with their own dataset? What errors/bugs could appear if the code were run in other devices/systems? What changes should be made to the code if it is run on a device with/without a GPU? How could the code be made more readable and easier to use for people who are interested in this paper? The latter, for example, could be achieved by naming variables appropriately and adding comments where necessary.
3. How do you recommend that people use the project repository you shared? Can they use the trained model you shared as is, or should they generate their own training datasets?
I would highly recommend this project repository to people interested in CEST analysis using deep learning. Although they cannot use the trained model directly if their experimental setup is different (in terms of CEST parameters, B0, subject type, etc.), the shared code is applicable to different CEST studies and can easily be implemented for training their own CEST data. People only need to prepare their input data in an appropriate format and feed them into the shared code. Once the network is trained, it can be used to quickly process CEST data in their project.
4. Are there any other reproducible research habits that you didn’t use for this paper but might be interested in trying in the future?
As regards data sharing, I recently noticed that Zenodo is another general-purpose open repository that allows researchers to deposit research papers, datasets, software, reports, and any other research-related digital artifacts. Moreover, a persistent DOI is assigned to each submission, which makes the stored items easy to cite. I am interested in trying this open repository in the future.




